*New Ph.D student in Team Phlow*
We are very happy to welcome Khadija Rombi as a CEPLAS-funded Ph.D student to our team! Khadi will be studying the communication systems required for logistics between source and sink tissues. Welcome to Team Phlow Khadi!
*New intern student in Team Phlow*
We are very excited to have Sonya Grozdani as intern student in our team! Sonya will be investigating methods to transdifferentiate nonvascular cells into phloem cell types! Welcome to Team Phlow Sonya!
1) Phloem development
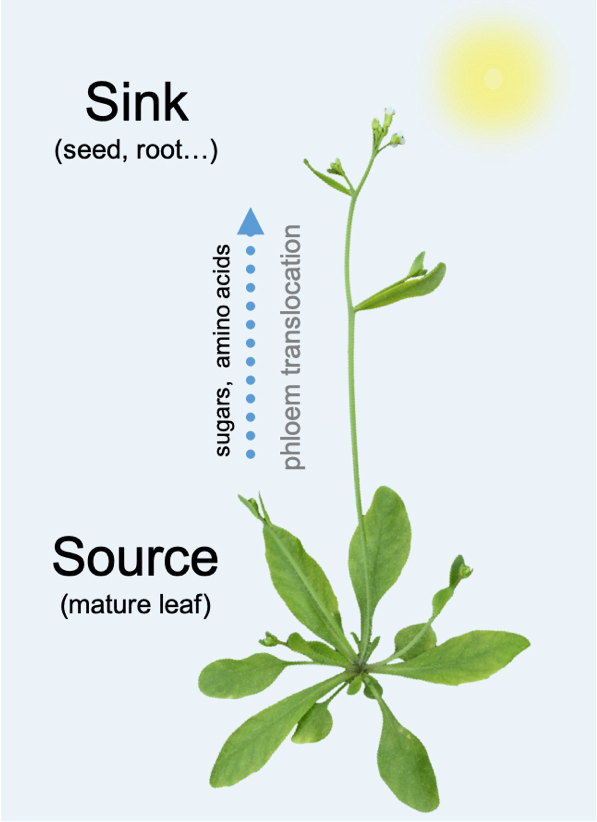
The distribution of assimilates between the source and sink organs is one of the most critical processes that must be properly coordinated in plants. Phloem loading, the process in which assimilates are loaded into the sieve element (SE), is the starting point for the translocation of assimilates across the whole plant. Our group aims to map the process of source-to-sink translocation of assimilates and understand diverse transport processes and the development of distinct phloem cell types.
We ask the following questions:
- Which transport processes are represented in the phloem?
- Which factors are involved in the development of the phloem?
- Can we identify strategies to enhance the source-sink translocation of sugars and amino acids?
We use a wide range of approaches to answer these questions, including single-cell RNA sequencing, translational profiling, CRISPR/Cas9-based gene editing, and transport assays. We anticipate that the knowledge gained from our project will provide a basis for assimilate allocation pathways and provide strategies to improve plant fitness and yield.
2) Matryoshka biosensors for monitoring and manipulating carbohydrate and gibberellin (GA) dynamics
As a proof of concept, we are establishing in vivo and in situ measurements of sugar dynamics in Arabidopsis leaves using a novel set of Matryoshka sugar sensors. Following successful implementation, the sugar sensor will be used for measuring sugar dynamics in Arabidopsis sink tissues. In parallel, we are also developing Matryoshka-type GA sensors to establish novel tools for controlling GA signaling networks. Ultimately, we aim to implement these tools in barley meristems to evaluate the delivery, synthesis, and turnover of GA in the meristem in different physiological conditions.
3) dCAS9-TurboID for identification of sugar transporter regulators
Cell-type specific expression of nutrient transporters is critical for resource allocation in plants. Exploring the chromatin composition of sugar transporters promoter region could possibly identify key transcriptional regulators of sugar allocation. By expressing promiscuous labelling enzyme fused with catalytically dead RNA-guided nuclease Cas9 (dCAS9) in mammalian cells, researchers were able to analyze the chromatin composition around the guide RNA targeting site by mass spectrometry. We are innovating a similar approach and developing plants harbouring dCas9 fused with TurboID-based proximity-labelling to identify potential regulators involved in sugar allocation during various plant growth conditions.

Arbeitsgruppenleiterin
Etage/Raum: 00.114