HOME | TEAM | FORSCHUNG | LEHRE | NEWS | PUBLIKATIONEN | STELLENANGEBOTE | RESOURCEN
Willkommen beim Institut für Molekulare Physiologie
Das Institut für Molekulare Physiologie untersucht die molekularen Grundlagen pflanzlicher Lebensprozesse. Im Fokus stehen Mechanismen, mit denen Pflanzen auf Umweltreize reagieren, Nährstoffe aufnehmen, Krankheiten widerstehen und ihr Wachstum regulieren. Ziel ist es, ein vertieftes Verständnis dieser Prozesse zu gewinnen und so zur Entwicklung nachhaltiger landwirtschaftlicher Strategien und widerstandsfähiger Pflanzensorten beizutragen.

Wolf B. Frommer
Institutsleitung
+49 211 81 12779
E-Mail senden
Hiroko Saito
+49 211 81-14826
E-Mail senden
Sabine Ahrens
+49 211 81-11575
E-Mail senden
News
Nora Zöllner verteidigt erfolgreich ihre Doktorarbeit mit Auszeichnung
Unsere Doktorandin Nora Zöllner hat ihre Dissertation mit dem Titel „Characterization of the infection process in rice bacterial blight by Xanthomonas oryzae pv. oryzae“ erfolgreich verteidigt und mit der Gesamtnote summa cum laude abgeschlossen, einer besonderen Auszeichnung für herausragende wissenschaftliche Leistungen.
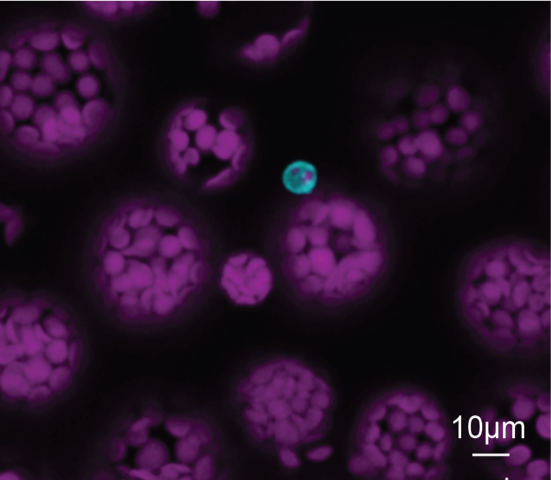
In ihrer Arbeit untersuchte sie die bakterielle Weißblättrigkeit in Reis, eine der weltweit bedeutendsten Pflanzenkrankheiten. Dabei konnte sie zeigen, wie das Pathogen Xanthomonas oryzae pv. oryzae gezielt den pflanzlichen Stoffwechsel manipuliert, um sich Zugang zu Zuckerressourcen zu verschaffen. Ein zentraler Beitrag ihrer Forschung ist die Identifizierung eines konservierten sux-Genclusters, das die Aufnahme und Verwertung von Saccharose ermöglicht. Zudem entwickelte Nora innovative Biosensoren, mit denen sich Zuckerflüsse in der Interaktion zwischen Wirt und Pathogen präzise verfolgen lassen. Ihre Ergebnisse liefern wichtige neue Einblicke in den Infektionsprozess und eröffnen vielversprechende Ansätze für die Entwicklung resistenter Nutzpflanzen.
Wir gratulieren Nora herzlich zu diesem herausragenden Erfolg und wünschen ihr für ihren weiteren Weg alles Gute!
Förderung für Forschung zu optimierten pflanzlichen Mikrobiomen
Dr. Eliza Loo hat eine Förderung aus dem Klaus Tschira Boost Fund für ihr Forschungsprojekt zu pflanzlichen Mikrobiomen erhalten.
Pflanzen leben in enger Wechselwirkung mit komplexen mikrobiellen Gemeinschaften, ihrem Mikrobiom, das entscheidend zu Nährstoffversorgung, Wachstum und Krankheitsresistenz beiträgt. Diese natürlichen Interaktionen gezielt nutzbar zu machen, etwa durch sogenannte Bioinokulanzien, gilt als vielversprechender Ansatz für eine nachhaltigere Landwirtschaft. Allerdings ist bislang noch unzureichend verstanden, wie sich solche nützlichen Mikroorganismen in bestehende mikrobielle Gemeinschaften integrieren.
Das geförderte Projekt „Predictive Rhizobiome Networks for Functional Microbiome Engineering“ setzt genau hier an. Ziel ist es, wurzelassoziierte Mikrobiome systematisch zu kartieren und ihre metabolischen Aktivitäten zu analysieren. Auf dieser Grundlage sollen Vorhersagemodelle entwickelt werden, die zeigen, welche Mikroorganismen erfolgreich in bestehende Gemeinschaften integriert werden können und somit das Pflanzenwachstum optimal unterstützen.
Langfristig soll die Forschung dazu beitragen, gezielt leistungsfähige mikrobielle Konsortien zu identifizieren und einzusetzen, um Nutzpflanzen robuster und produktiver zu machen.
Weitere Informationen im Originalartikel: Zum vollständigen Beitrag
Pflanzen stärken, Zukunft sichern
Eine öffentliche Veranstaltung beleuchtet Innovationen in der Pflanzenforschung
Das Projekt „Healthy Crops“ veranstaltete gemeinsam mit der Bürgeruniversität der Heinrich-Heine-Universität Düsseldorf im Haus der Universität die öffentliche Veranstaltung „Pflanzen stärken, Zukunft sichern – Forschung für gesunde Ernten“. Ziel war es, aktuelle Herausforderungen und Innovationen der Pflanzenwissenschaft durch eine Kombination aus Vorträgen und interaktiven Formaten einem breiten Publikum näherzubringen. Den Auftakt bildete ein Vortrag von Dr. Marcel Buchholzer zur globalen Ernährungssicherheit, in dem Reis als Modellsystem diente, um das Zusammenspiel von biotischem Stress, Klimawandel und Bevölkerungswachstum zu veranschaulichen. Dabei wurden Strategien zur Sicherung und Steigerung von Erträgen vorgestellt, von klassischer Züchtung bis hin zur Genom-Editierung. In einem zweiten Vortrag erläuterte Prof. Wolf Frommer die zentrale Rolle von Mutationen als Motor von Evolution und Vielfalt und ordnete deren Nutzung in der Pflanzenzüchtung ein. Im Anschluss wurde das Projekt „Healthy Crops“ als Lösungsansatz für biotische Herausforderungen präsentiert, wobei entwickelte Pflanzenlinien und deren Bedeutung für eine nachhaltige Landwirtschaft im Fokus standen.
Der interaktive Teil „Plant Research Hands-On“ ermöglichte den Teilnehmenden einen direkten Zugang zur Pflanzenforschung. Neben mikroskopischen Untersuchungen verschiedener Reissorten konnten Besucher ihr Wissen in Quizformaten testen und sich anhand einer Weltkarte über die globale Arbeit des Projekts informieren. Weitere Stationen veranschaulichten die Vielfalt von Brassica-Kulturen sowie die Grundlagen der Genom-Editierung. Die Veranstaltung verband erfolgreich wissenschaftliche Tiefe mit verständlicher Vermittlung und förderte den Dialog zwischen Forschung und Öffentlichkeit. Sie unterstrich die Bedeutung interdisziplinärer Ansätze zur Bewältigung globaler Herausforderungen der Ernährungssicherung.
Laura Redzich hat ihre Doktorarbeit erfolgreich verteidigt
Wir gratulieren unserer Doktorandin Laura Redzich herzlich zur erfolgreichen Verteidigung ihrer Doktorarbeit mit dem Titel “Spatiotemporal investigation of Xanthomonas oryzae pv. oryzae infection trajectories and the functional impact of SWEET promoter editing”.
In ihrer Arbeit untersuchte Laura die räumlich zeitliche Dynamik der Infektion von Reis durch ein weitverbreitetes Reis-Pathogen und analysierte den funktionellen Einfluss der gezielten Editierung von SWEET Promotoren auf die Krankheitsentwicklung. Ihre Ergebnisse liefern wertvolle neue Einblicke in die molekularen Interaktionen zwischen Pathogen und Wirt und tragen wesentlich zur Entwicklung innovativer Strategien für eine nachhaltige Krankheitsresistenz in Reis bei.
Wir danken Laura für ihr großes Engagement und ihre hervorragende wissenschaftliche Arbeit und wünschen ihr für ihren weiteren beruflichen und persönlichen Weg alles Gute und viel Erfolg.
Genome editierte Reissorte Kamala zeigt deutliche Ertragssteigerung in indischen Feldversuchen
Ergebnisse aus den landesweiten indischen AICRIP (All India Coordinated Research Project on Rice) Feldversuchen zeigen, dass die genome editierte Reissorte Kamala im Durchschnitt eine Ertragssteigerung von 19 Prozent gegenüber etablierten Vergleichssorten erreicht. Kamala basiert auf einer elitären Reissorte und wurde durch die gezielte Feinabstimmung der Aktivität eines Cytokinin Oxidase Gens mittels Genomeditierung entwickelt.
Die erhöhte Ertragsleistung wurde konsistent unter praxisnahen Feldbedingungen beobachtet und unterstreicht das Potenzial präziser genome editierter Ansätze zur nachhaltigen Steigerung der landwirtschaftlichen Produktivität. Die wissenschaftlichen Ergebnisse wurden als Preprint auf bioRxiv veröffentlicht (Link zur Publikation).
Die Entwicklung von Kamala erfolgte in enger Zusammenarbeit mit Satendra Mangrauthia und indischen Partnerinstitutionen. Die Linie wurde inzwischen offiziell als neue Reissorte in Indien registriert und kann auf Grundlage der geltenden Regularien der indischen Regierung prinzipiell für den Anbau freigegeben werden, da sie keine transgenen Sequenzen enthält.
Die Registrierung von Kamala wurde öffentlich von der Organisation The Coalition for a GM Free India kritisiert. In diesem Zusammenhang hat das Indian Council of Agricultural Research (ICAR) eine offizielle Stellungnahme veröffentlicht, die die wissenschaftliche und regulatorische Einordnung der Sorte erläutert (Link zur Stellungnahme).
Unsere Nationalitäten